By Bernd Pulverer, EMBO
As preprint posting takes hold in the biosciences community, we need both quality control and curation to ensure we share results in a reproducible and discoverable manner
The EC has taken the bold step – at least on paper – to proclaim a Europe that is by 2020 to be ‘Open Innovation’, ‘Open Science’ and, thankfully, ‘Open to the World’. The Open Science piece includes an ongoing project ‘The Open Science cloud’ as well as a dedicated online publishing/preprint platform – details pending at the time of writing. The promise of Open Science is of course no less than to dramatically increase the efficiency of the scientific process. Given the speed of these developments, now is the time to face up to a key issue in the life sciences: the question of trust and quality of data shared by Open Science mechanisms. Clearly, a 24/7 release of raw data by all labs will rapidly clog up even the most powerful repositories, while providing only limited benefit to the community. Without the provision of stable, structured databases and repositories that are curated and quality-controlled, we risk sinking in a swamp of data that will be largely ignored.
Clearly, preprints ought to play a key part in any Open Science agenda as a bridge between data and research papers. Preprints are growing exponentially and are poised to be accepted as a community standard in the biosciences. Now is the time to establish mechanisms that ensure we select for rigorous results that are associated with sufficient metadata to render them reproducible and discoverable. In the physical sciences, arXiv has evolved a large and proactive community of volunteers that screen thousands of prospective posts each month. Plans for the semi-automation of these processes are pending. bioRXiv has taken a similar approach with a collegium of volunteers who do the most basic of assessments: ‘does the preprint look like a scientific paper’. In my view, while laudable and necessary, this will not be sufficient in the long run. To be sure, key to the success of preprints is ease and speed of submission. As the volume is increasing exponentially, we need to find scalable ways to integrate quality control steps that reinforce the rigor and utility of the science shared, while posing minimal friction for researchers.
Reproducibility
We need to ensure preprints include sufficient metadata to render the experimental data reproducible by others. This is an opportunity to go beyond what the average scientific journal can muster by fortifying emerging authoring tools with templated methods sections that allow for the reporting of materials and methods in a structured, machine-readable way (including, where available, reagent bar coding and link-outs to resources that report detailed protocols and allow the unequivocal identification of specific reagents).
Data Discoverability & connecting experiments
The data presented in figures and tables represent the core of the scientific evidence in a manuscript. As a consequence, the level of confidence a reader has in non-peer reviewed scientific work will be reinforced if preprints can be linked to each other and to published research papers via the experimental results shown in figures. For example, the EMBO SourceData platform converts figures and their legends into searchable machine-readable metadata describing the design of the reported experiments (sourcedata.embo.org). Interlinking the data presented in figures and preprints helps users to place a given preprint in the context of related data. This facilitates browsing of related content and also helps readers to determine the uniqueness, the degree of coherence and of replication of the reported experiments.
At the same time figures that are carefully curated by human or machine renders them directly discoverable by data-directed search technology. It will be crucial to enable researchers to interrogate the preprint knowledge base for specific experiments including the associated data, materials and methods
Quality
The assessment of the quality of preprints can happen at three levels:
- Are the experiments reported in a way that allows their interpretation and replication by others? Is the preprint marred by any problems related to research integrity or ethics (such as image manipulation, or sub-par statistics)?
- Are the experiments carried out in a robust manner that allows meaningful interpretation?
- Are the experiments reported valuable to the research community. Do they warrant preservation, curation and dissemination?
The current assessment of preprints does not cover any of these levels, at least formally. Indeed, I would argue we do not need to assess preprints at all for their semblance to a classical research paper and should be open to share smaller units of reproducible, fully described experimentation without the need for long free text introduction and conclusions. We need to decide which of these assessment levels should be implemented, and which would be desirable, assuming sufficient support and community engagement. Ideally, we will find a scalable way to systematically apply all three levels, but who would take on these responsibilities?
Who?
In my view, curation and quality control of the data can be executed optimally as an integrated workflow by quality checkers, be they editors, curators or researchers, who are assisted by state-of-the art computer automation (#1 above). Indeed, once the process is clearly defined, a high level of automation can be achieved based on the application of AI-trained on the human-curated datasets. Automatic curation will be subject to professional quality checks and author-based validation through user-friendly interfaces with simple yes/no functionality (see sourcedata.embo.org).
Screening for work that is meaningful and valuable to the community (#2 & 3 above) will have to be done by knowledgeable experts. This might in principle either include the established pool of senior academics who review papers – as proposed elsewhere. However, in my view we ought to tap into the pool of more junior researchers to avoid exacerbating the peer review bottleneck. There is a vast and highly capable community of experienced postdocs who would be very well placed to carry out these tasks. They would be incentivized by including this activity as a quantifiable output relevant for research assessment, or by direct financial compensation at a level that would be affordable even at high volumes (for example, 50$/manuscript). A second group might be retired academics interested in staying engaged with the community.
Integration with Journal workflows
Journals apply peer review and editorial assessment to submitted manuscripts. More progressive journals have started to apply more or less complex additional screening processes to complement peer-review. At the EMBO publications this involves in-house screening for statistics and image aberrations as well as curation. The curation step will be at least partially automated in the future and culminates in an easy to use author validation interface (for more information on the EMBO Source Data paradigm see sourcedata.embo.org ). It is imperative to avoid redundancy in such screening processes as they require significant overheads and incur delays. Many in the community are concerned about scooping, and many preprints that are posted are simultaneously submitted for publication – something journals have started to facilitate with co-submission/preprint posting functionality in their online systems. We therefore hope that the quality control and curation exercise can be applied to preprints by executing it at this point of intersection of preprints and journals.
However, for cost reasons many journals will not apply such screening processes in their workflows and to be sure not all preprints need be destined for journal publication in the first place. We should ultimately apply the screens outlined here to all preprints systematically. Once the issues of quality control, reproducibility and discoverability are addressed in preprints, they will be taken over to the published journal paper, as it represents no extra costs for the journal and the journal will not want to be less useful than the preprint to the reader. In other words, a central quality-controlled preprint server can play an important role in improving the literature at many journals. Funders could include the expenses for quality control and curation in their research awards.
Conclusions
As others have suggested on this blog, full peer review may also be applied to preprints and they may be subjects for journal clubs, as Martin Chalfie suggested at ICCB2018 this week – he reported that his lab has switched with considerable enthusiasm to doing journal club on preprints, and posts the results to the authors (he credited Francis Collins with the concept). The Company of Biologists will soon launch its own community preprint commenting platform ‘preLights’. These are all important and valuable post-posting initiatives, but what we need first and foremost is a pre-posting filtering mechanism.
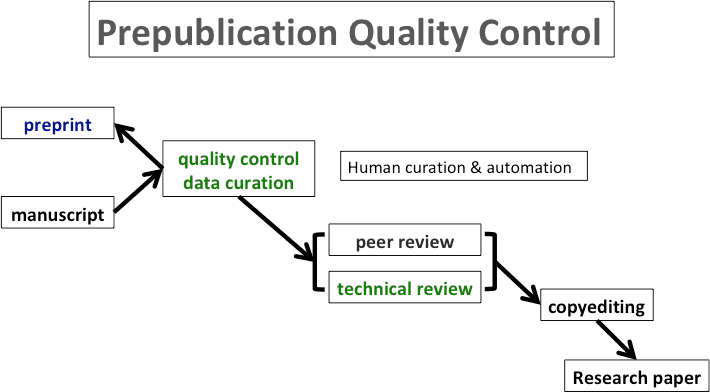
Fig. 1. Quality control and data curation steps applied prior to preprint posting, integrated with the journal submission system.
Ancknowledgement: with thanks to Thomas Lemberger and Maria Leptin for suggestions.