The growth of preprints in the life sciences has amplified earlier concerns about the challenges of keeping abreast of the latest research findings. Researchers need to keep up to date not only with the most recent publications in journals but also with the latest scholarly work posted on preprint servers. Three quarters of the respondents to the #biopreprints2020 survey last year noted information overload as very or somewhat concerning in the context of preprints. The facilitators of one of the breakout session at the #FeedbackASAP meeting outlined proposals to help researchers find the latest relevant preprints and also to attract preprint reviewers.
Finding relevant preprints
Christine Ferguson and Martin Fenner outlined their proposal to develop ways for researchers to find preprints relevant to their research immediately after the preprints appear. They propose an automated system that would identify preprints posted in the previous few days that had received attention via Twitter (i.e. based on the preprint receiving a minimal number of tweets).
During the discussion, the session attendees mentioned a number of currently-available tools that collect reactions and attention on preprints and/or allow researchers to discover the latest preprints:
- CrossRef collects Event Data for individual preprints, including social media mentions, Hypothes.is annotations and more.
- The Rxivist.org tool allows searching for bioRxiv and medRxiv preprints based on Twitter activity.
- The search.bioPreprint tool developed by the University of PIttsburgh Medical Library allows searching preprints from different servers based on keywords or topics.
- bioRxiv provides search options based on discipline and also has a dashboard that collects reactions and reviews on individual preprints, including Twitter comments.
- EMBO has developed the Early Evidence Base platform which allows searching for refereed preprints.
- Google Scholar indexes preprints and provides some filtering tools.
The attendees raised some questions about the use of Twitter as a filter and the risks for a metric based on tweets. How can we account for the risk of social media users gaming the system by artificially boosting attention on Twitter? How can we normalize for the fact that methods papers tend to receive more attention? Is there a risk that this system will be focused on papers from high-income countries that already receive a disproportionate share of attention?
Christine and Martin noted that their proposal involves a direct weekly email to researchers as the channel for communication, allowing those who sign up to relevant subject areas to receive a list of preprints without them having to individually mine different tools. Their proposal does not sort or rank papers by the number of tweets, but rather requires a minimum threshold of Twitter activity for inclusion. They recognized that there may be a number of biases associated with attention to preprints, some we are aware of and some which we still may not know about, but they propose their approach as a starting point which can be developed further.
The group suggested setting up a form to which researchers could sign up to signal interest in receiving the proposed newsletters. To this end, those interested in using and feeding back on the early prototype, can sign up via this link: https://front-matter.io/newsletter, and will receive weekly lists of preprints in specific subject areas. The feedback will be used to fine-tune the newsletters going forwards.
Finding reviewers
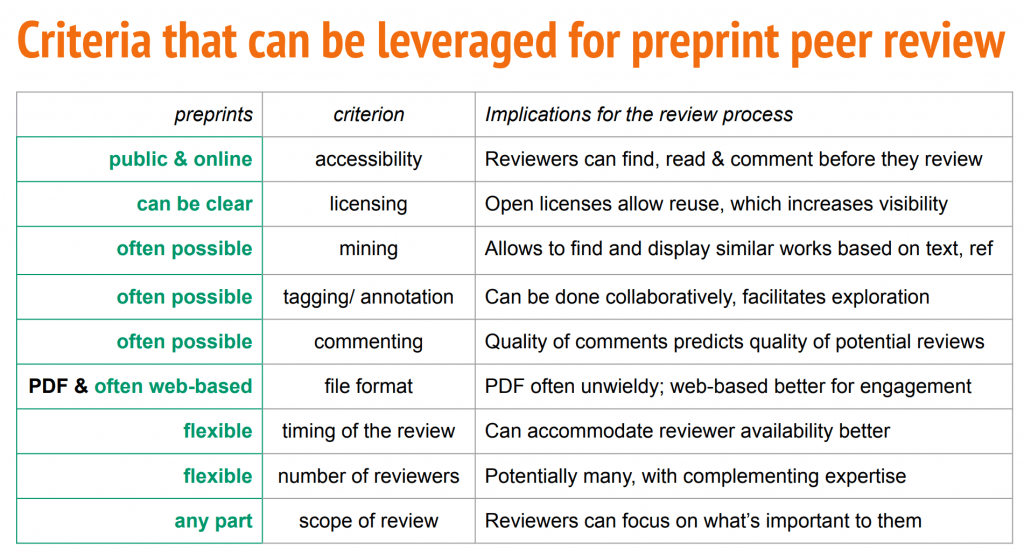
In the last part of the session, Daniel Mietchen outlined the differences between preprint and journal review and how some of the advantages of preprint review can be leveraged to find reviewers.
Daniel proposed some recommendations for how preprint servers could attract preprint reviews:
- Use open licenses for preprints and formats that allow reuse.
- Provide mechanisms to flag which aspects of a preprint are in most need of a review – this could be done by authors themselves or by readers who indicate whether e.g. the statistics or a specific methodology should be evaluated by another expert.
- Make it easy to annotate preprints by having HTML versions and enabling Hypothes.is.
- Use non-tracking tools for commenting.
- Make it easy to subscribe to content alerts using specific filters, e.g. based on mentions to specific concepts (cell line, model organism, author, institution) in sections of the paper or to lists of preprints/papers with high similarity to a given piece of text (similar to the JANE tool to search PubMed).
Christine (ferguson.cav@gmail.com), Martin (martin@front-matter.io) and Daniel welcome comments and feedback on their proposals, feel free to comment on this blog post or contact them directly.